About
What does this tool do?
PET2BIDS accepts PET imaging and blood data as inputs (e.g. dicom, ecat, spreadsheets) and delivers BIDS fomatted outputs (e.g. nifti, json, and tsv files). See the below diagram:
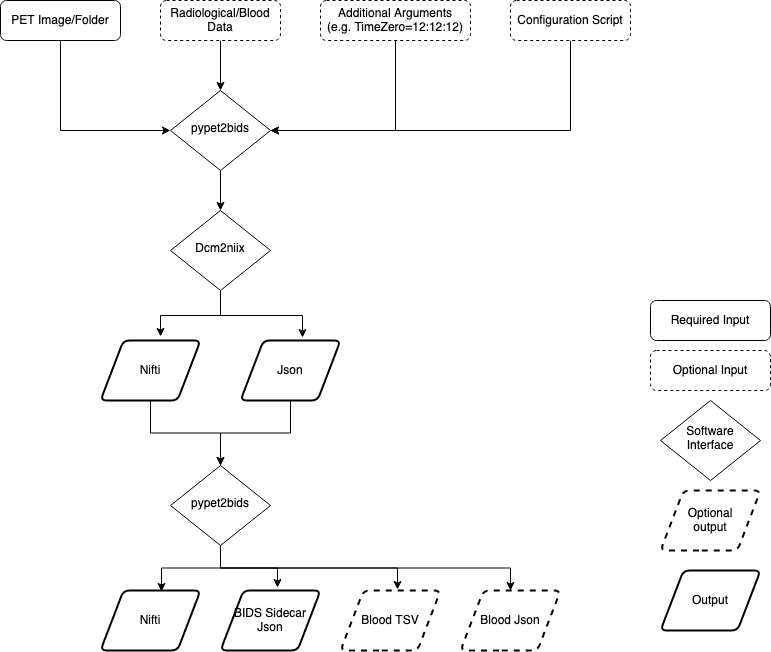
It looks like you’re just wrapping dcm2niix, why not just use it?
That’s a correct assessment, however:
dcm2niix only handles the conversion from dicom to nii and knows roughly what BIDS is, but the json file produced is not fully compliant with the standard
dicom data often doesn’t have radioactivity information (e.g. injected dose) and that’s why you need to provide it
conversion of blood data requires dedicated scripts and as dcm2niix has no idea what a spreadsheet is (which is implicit in the name dicom 2 niix)
Where is the User Interface?
The PET2BIDS library does not have an interface. We are however working with BIDSCoin and ezBIDS to integrate our library into their GUI. You can however use the code as such.
What is the command line for matlab?
After setting up the library in your matlab path only 3 commands are needed: meta = get_pet_metadata(varargin) followed by either ecat2nii(file, meta) or dcm2niix4pet(dcmfolder,meta,'o','mynewfolder'). For a comprehensive list of available commands/methods see matlab.
What is the command line for Python or better, is there or command line interface?
Yes there is a CLI and no you don’t need to learn Python at all, but you’re more than able to use this tool as a Python library. For more information on usage individual methods or modules refer to pypet2bids.
How do I use this tool?
Please refer to the Usage page for details on how to go about using this library or the quickstart page for a small walk-through. Additionally, if you want to truly see how this software works on real data, take a look at our CI on Github as a further demonstration of running this software.
This tool doesn’t work/this is really hard….
Reshaping PET data into BIDS can often be difficult, but it’s the goal of this library and it’s developers to make the process easier for you the user. If you are struggling with using the tools (or the tools are struggling to work with your data) reach out us via our Issues Page or through our Website/Email.